Todd Hollon
Bio
I am an associate tenured professor and the Novello & Quinones-Hinojosa Research Professor at the University of Michigan. I am the principal investigator of the Machine Learning in Neurosurgery (MLiNS) Lab, where our research focuses on developing machine intelligence that understands human health and disease, especially related to the nervous system. We aim to discover better data streams, model architectures, inductive biases, and learning objectives for medical AI. Our technical contributions include improved visual self-supervision, hierarchical and multimodal representation learning, and medical foundation modeling.
We are actively hiring phd students and postdocs in medical AI. Projects can cover any of the research themes below. Please email me directly at tocho [at] med.umich.edu if interested.
Photos and Bios of MLiNS Team
The team, the team, the team.
MLiNS News
- May 2026 - FastGlioma, published in Nature, is featured by UM Look to Michigan campaign.
- Apr 2026 - Xinhai Hou’s paper, CodeV, is accepted at CVPR as Oral Paper (Top 1%).
- Apr 2026 - Samir Harake wins UM Khan Neurosurgery Award for best medical student.
- Apr 2026 - Rush Joshi wins Best Clinical Absract at 2026 Neurosurgery Symposium.
- Feb 2026 - Prima is published in Nature Biomedical Engineering.
- Feb 2026 - Yiwei Lyu’s paper, ItemizedCLIP, is accepted at CVPR.
- Dec 2025 - MLiNS lab’s featured in The Detroit News by UM president Domenico Grasso.
- Mar 2025 - Todd Hollon named the inaugural Joseph R. Novello, M.D. and Alfredo Quiñones-Hinojosa, M.D., Ph.D., Research Professor of Neurosurgery.
- More news here.
MLiNS Research Themes
- Intelligent Histology - intraoperative microscopy, computational pathology, brain tumor imaging
- AI-based Neuroimaging - AI for brain, spine, and peripheral nerve imaging
- Visual Intelligence - visual reasoning, visual representation learning, vision-language modeling
- Patient Forecasting - outcome and survival prediction
- Collaborative Neuro-Oncology - team science in brain tumor research
Selected Publications and Complete List
Biomedical research journals
-
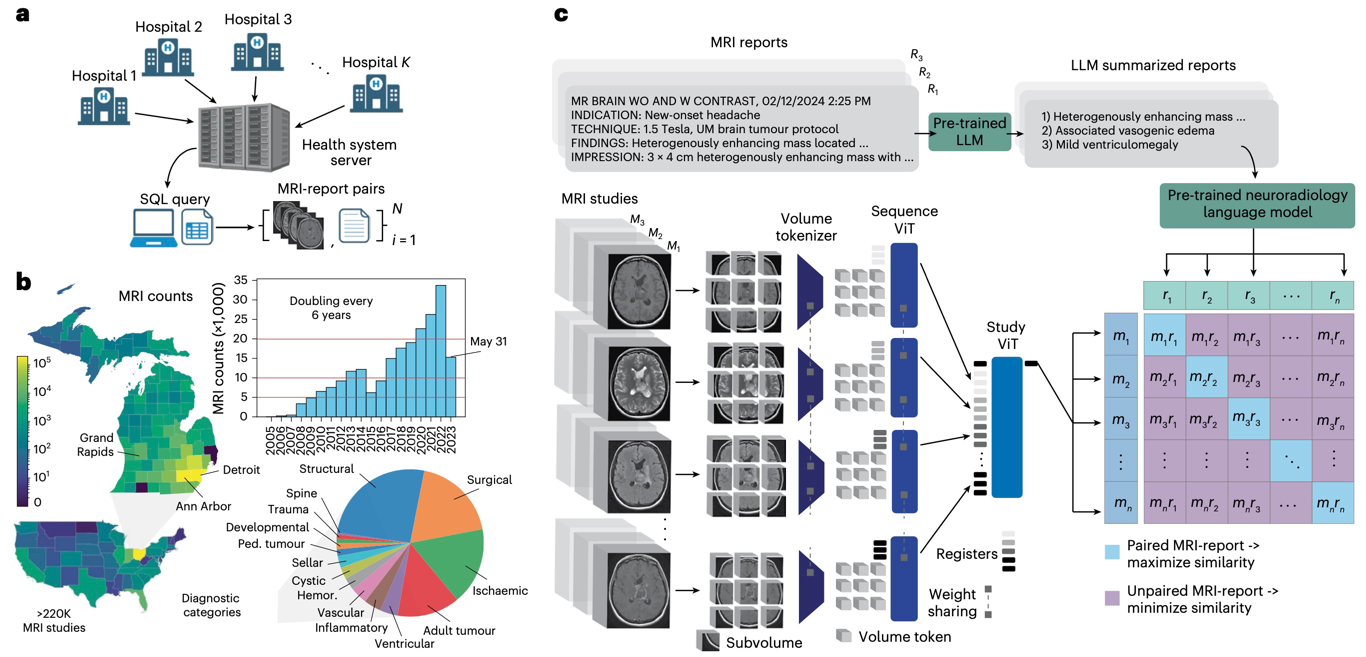
 NATURE BIOMEDICAL ENGINEERING · 2026
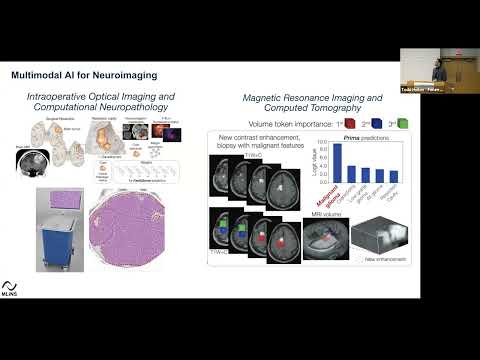
NATURE BIOMEDICAL ENGINEERING · 2026Introduces Prima, a visual foundation model for brain MRI trained at health-system scale for diagnosis, triage, and clinically grounded decision support.
-
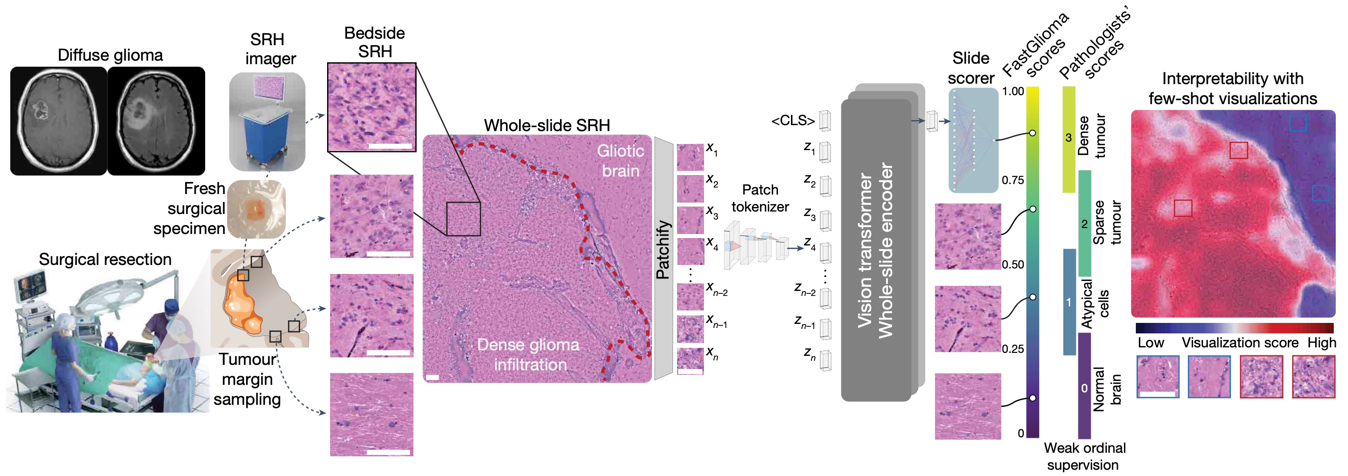
 NATURE · 2025
NATURE · 2025FastGlioma is a computational pathology model for real-time detection of glioma infiltration at the surgical margin, outperforming the current standard of care.
-
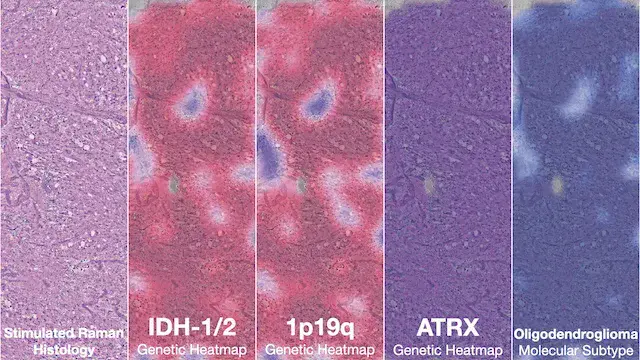
 NATURE MEDICINE · 2023
NATURE MEDICINE · 2023Diffuse gliomas are classified using the molecular features. DeepGlioma predicts the molecular genetics of brain tumors within minutes of biopsy, in the operating room, to better inform surgical goals.
-
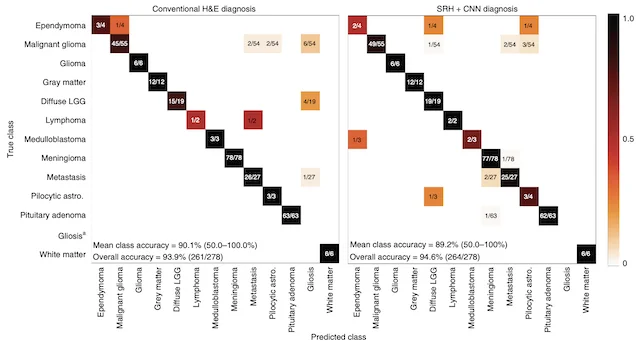
 NATURE MEDICINE · 2020
NATURE MEDICINE · 2020A deep learning workflow combining stimulated Raman histology with convolutional neural networks delivers near real-time intraoperative brain tumor diagnosis, matching pathologist accuracy while compressing turnaround from ~30 minutes to under 150 seconds.
Machine learning conferences and proceedings
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2026
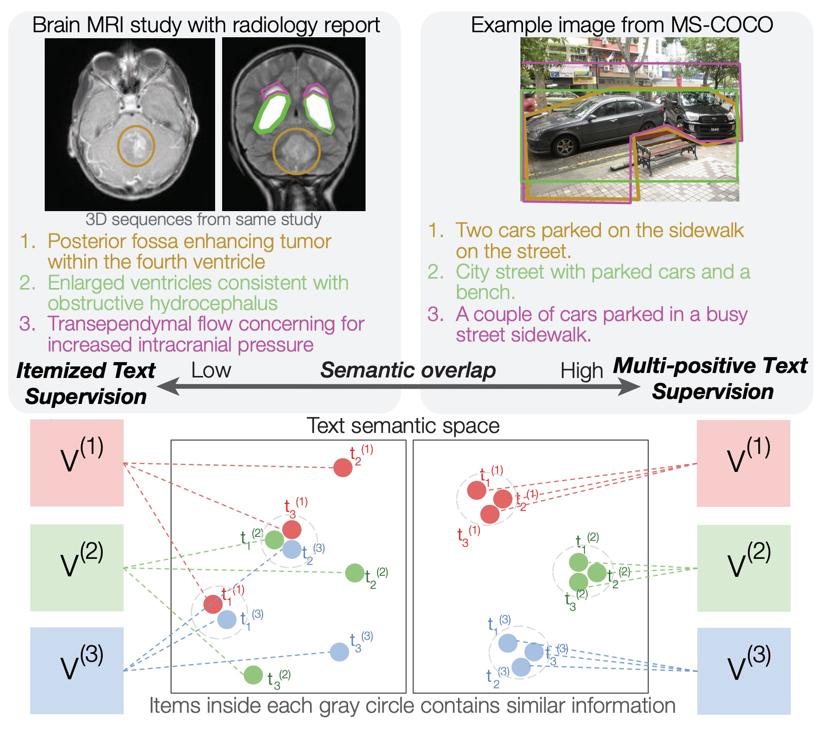
COMPUTER VISION AND PATTERN RECOGNITION · 2026Standard contrastive language-image pre-training can neglect objects in visual scenes. ItemizedCLIP forces models to learn and attend to all described items, resulting in better visual representations.
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2026
COMPUTER VISION AND PATTERN RECOGNITION · 2026Recent visual agents can score well while using image tools unfaithfully-e.g., cropping irrelevant regions or ignoring tool outputs. CodeV represents tools as executable Python code and trains with Tool-Aware Policy Optimization (TAPO), using process-level rewards on visual tool inputs and outputs to improve both accuracy and faithful tool use on search and broader multimodal benchmarks.
-
 TRANSACTIONS ON MACHINE LEARNING RESEARCH · 2026
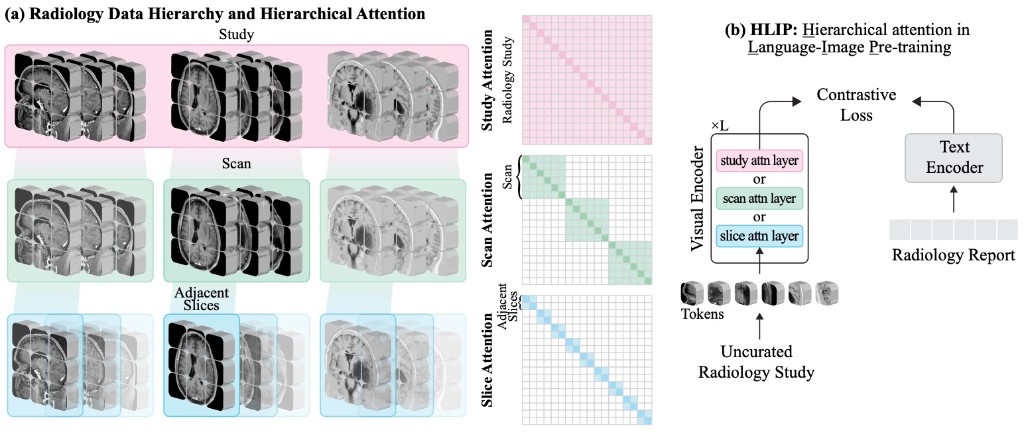
TRANSACTIONS ON MACHINE LEARNING RESEARCH · 2026Vision-language pre-training for volumetric MRI and CT is usually limited by radiologist-curated datasets. HLIP instead uses hierarchical attention over slice, scan, and study to pre-train on uncurated clinical data at scale, boosting performance on public brain MRI and head CT benchmarks.
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2023
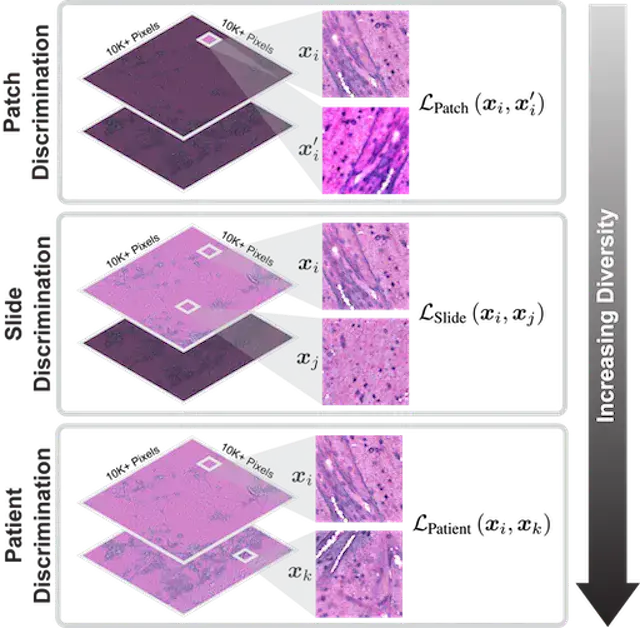
COMPUTER VISION AND PATTERN RECOGNITION · 2023HiDisc is a self-supervised learning method that leverages the inherent patient-slide-patch hierarchy of biomedical microscopy to learn stronger visual representations without explicit negative mining.
-
 NEURIPS DATASETS & BENCHMARKS · 2022
NEURIPS DATASETS & BENCHMARKS · 2022OpenSRH is the first public dataset of clinical stimulated Raman histology images from brain tumor patients, released alongside benchmarks to accelerate machine learning research for intraoperative brain tumor diagnosis.
New Preprints
-
 medRxiv 2026
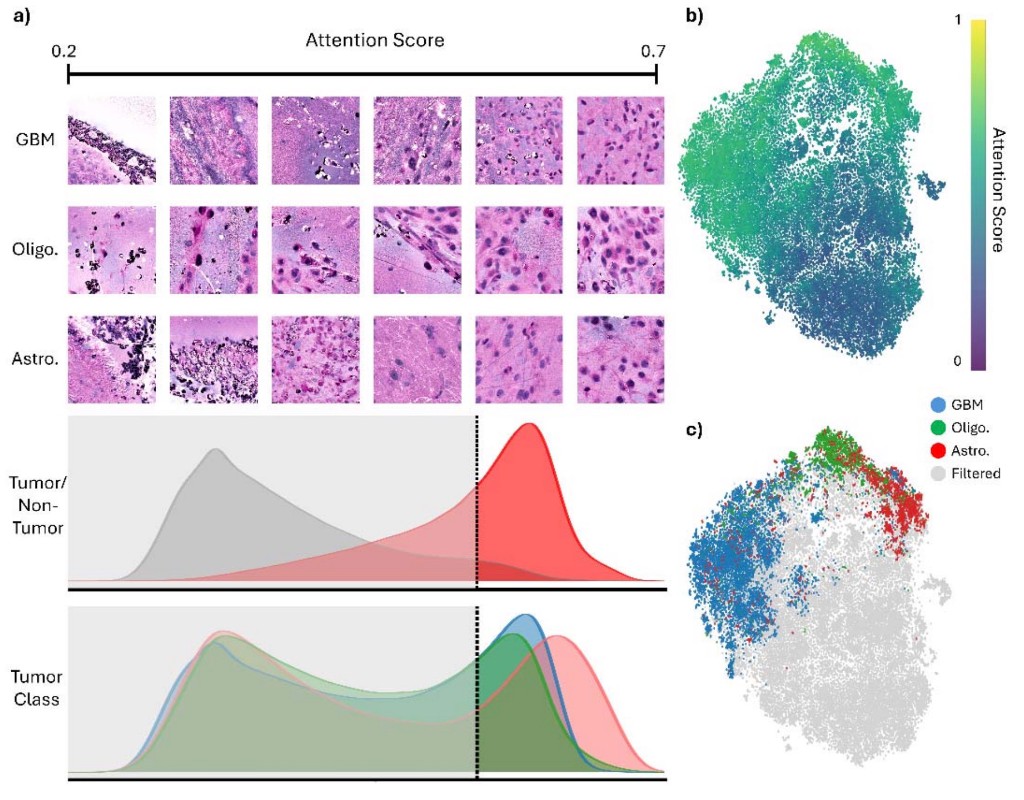
medRxiv 2026SIGNAL is a scalable model for rapid intraoperative molecular classification of gliomas from stimulated Raman histology, with attention-guided patch visualization that separates tumor from non-tumor tissue and distinguishes GBM, oligodendroglioma, and astrocytoma in real-world surgical settings.
-
 arXiv 2026
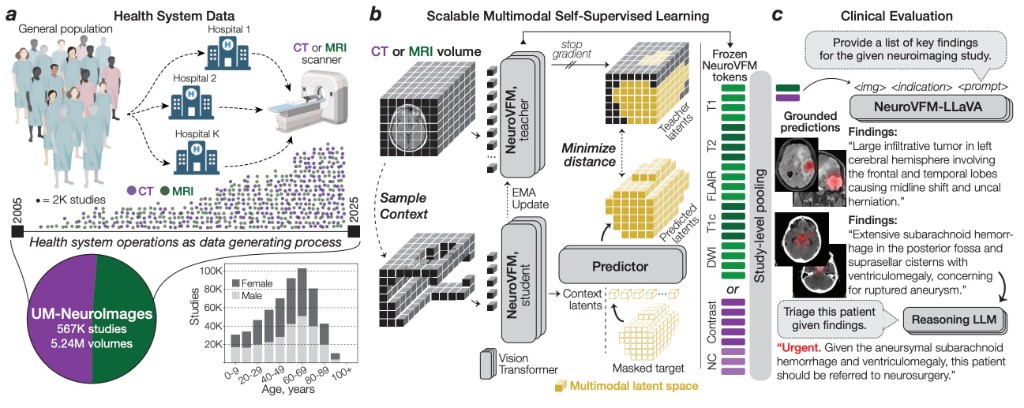
arXiv 2026NeuroVFM is a visual foundation model trained on 5.24M clinical MRI and CT volumes via health system learning, a paradigm that leverages uncurated data from routine care. Using a scalable volumetric joint-embedding predictive architecture, it delivers state-of-the-art radiologic diagnosis and report generation with interpretable visual grounding, surpassing frontier models in accuracy, triage, and expert preference while reducing hallucinations.
-
 medRxiv 2025
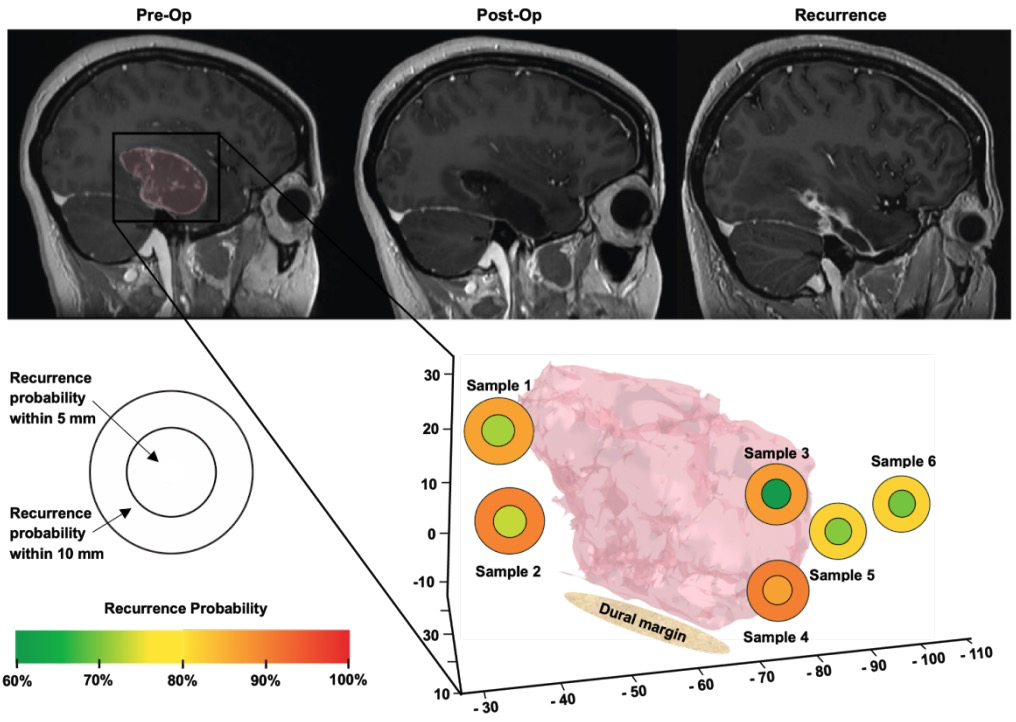
medRxiv 2025Using AI-quantified tumor infiltration from label-free optical microscopy of surgical margin tissue, combined with clinical, radiographic, and molecular features, a random forest model predicts sites of focal glioblastoma recurrence (validation AUC 80.3%). AI-derived infiltration was the strongest predictor, outperforming molecular features alone, pointing toward margin-guided precision adjuvant therapy for the highest-risk areas of disease.
MLiNS Videos and Podcasts
Improving the Results of Brain Surgery > Look to Michigan (YouTube)
I AM AI, NVIDIA GTC 2021, Official Intro (YouTube) (starts at 1:20)
More videos and podcasts here.
Support for MLiNS Lab
- NIH NCI R01
- NIH NINDS K12
- NIH NINDS F31
- Chan Zuckerberg Initiative
- Ian’s Friends Foundation (Phil and Cheryl Yagoda)
- UM Frankel Institute for Heart and Brain Health
- UM Scouts Program
- UM Department of Neurosurgery
- Scoliosis Research Society
- Cook Family Foundation
Please email me directly at tocho [at] med.umich.edu if you would like to support our effort. We are grateful for any contribution, big or small.