Todd Hollon
Visual Intelligence
Our visual world is more complex than human language. Language is discrete, sequential, and constructed. Our visual world is continuous, geometric, and discovered. A major open problem in machine intelligence is how best to model visual perception and reasoning. We believe that the major advances in large language models, while impressive, do not illuminate this problem. Our lab focuses on visual reperesentation learning broadly and how image data structures can inform visual learning. For example, we exploit inherent hierarchical structures in biomedical microscopy (HiDisc) or neuroimaging (HLIP) to better learn complete and grounded visual features. We also aim to unify visual self-supervision and languauge supervision (CLIPred, SimCLIP), which are generally treated as independent learning enviroments. Visual reasonsing enables AI agents to reason about images, performing actions on those images such as cropping, clipping, and resizing (CodeV).
Related publications
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2026
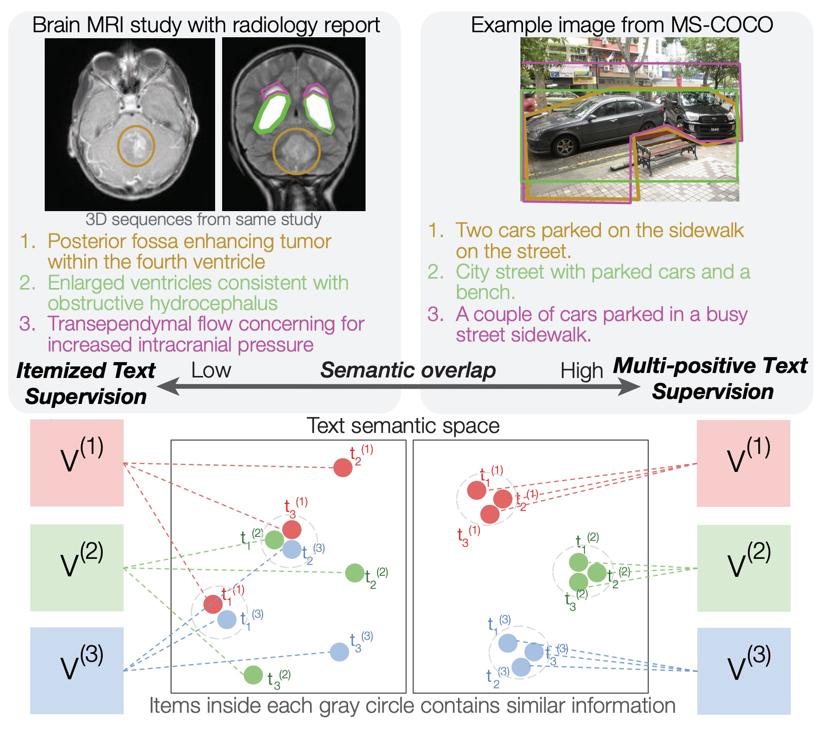
COMPUTER VISION AND PATTERN RECOGNITION · 2026Standard contrastive language-image pre-training can neglect objects in visual scenes. ItemizedCLIP forces models to learn and attend to all described items, resulting in better visual representations.
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2026
COMPUTER VISION AND PATTERN RECOGNITION · 2026Recent visual agents can score well while using image tools unfaithfully-e.g., cropping irrelevant regions or ignoring tool outputs. CodeV represents tools as executable Python code and trains with Tool-Aware Policy Optimization (TAPO), using process-level rewards on visual tool inputs and outputs to improve both accuracy and faithful tool use on search and broader multimodal benchmarks.
-
 NEURIPS UNIREPS WORKSHOP · 2025
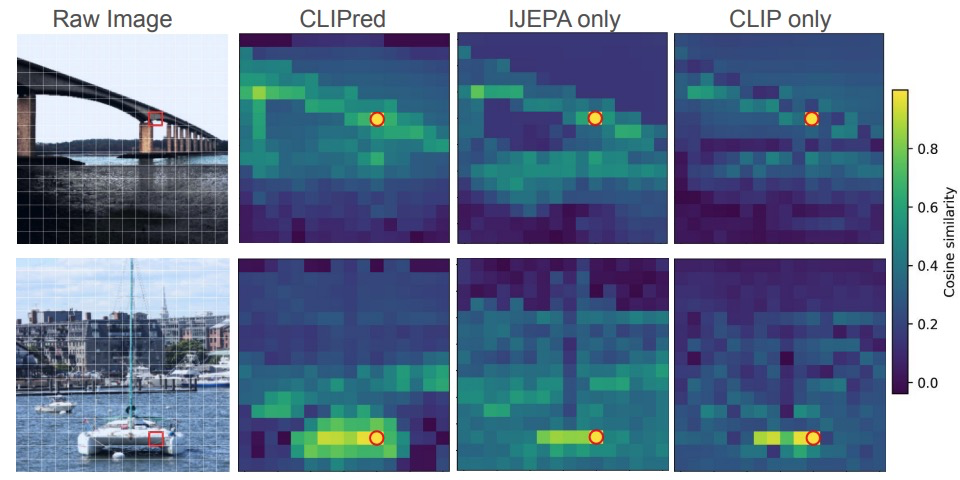
NEURIPS UNIREPS WORKSHOP · 2025CLIPred is a framework that jointly optimizes the I-JEPA self-supervision and CLIP language supervision objectives for visual representation learning, outperforming either alone and achieving better zero-shot transfer than DINOv2+CLIP at lower training cost.
-
 AAAI · 2025
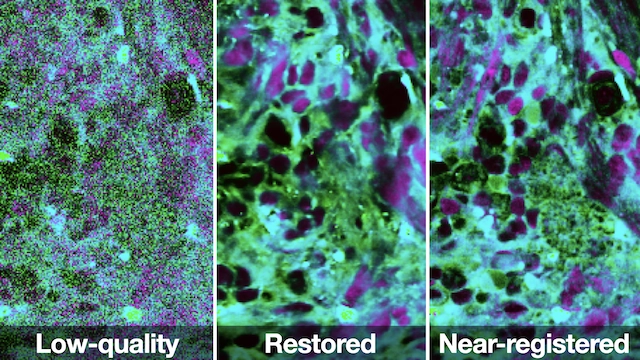
AAAI · 2025This paper introduces Restorative Step-Calibrated Diffusion (RSCD) for biomedical optical image restoration, improving reconstruction fidelity by adapting denoising dynamics to the characteristics of microscopy data.
-
 NEURIPS FITML WORKSHOP · 2024
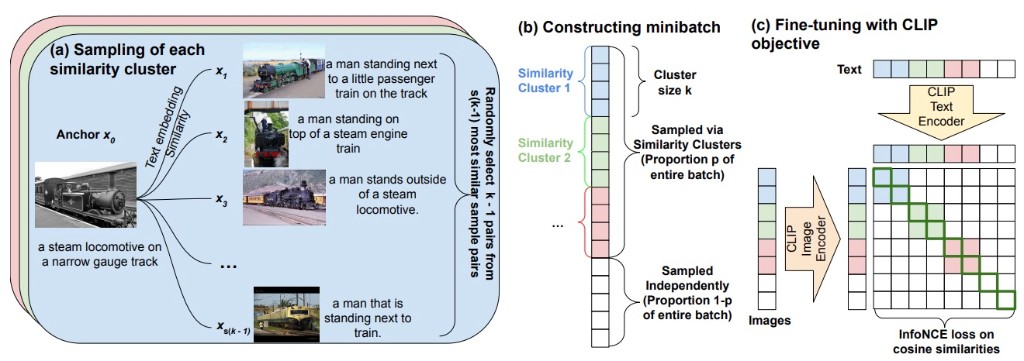
NEURIPS FITML WORKSHOP · 2024SimCLIP is a generalized framework for CLIP fine-tuning that constructs minibatches containing clusters of similar image-text pairs to produce harder in-batch negatives, improving downstream performance over standard CLIP fine-tuning without hand-crafted hard negative captions.
-
 CVPR WORKSHOP · 2024
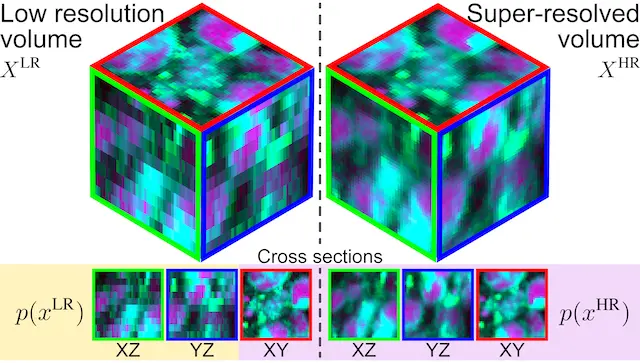
CVPR WORKSHOP · 2024This work proposes Masked Slice Diffusion for Super-Resolution (MSDSR), a strategy for volumetric biomedical super-resolution trained with only 2D supervision, enabling high-quality 3D reconstruction when fully paired 3D labels are scarce.
-
 ARXIV · 2024
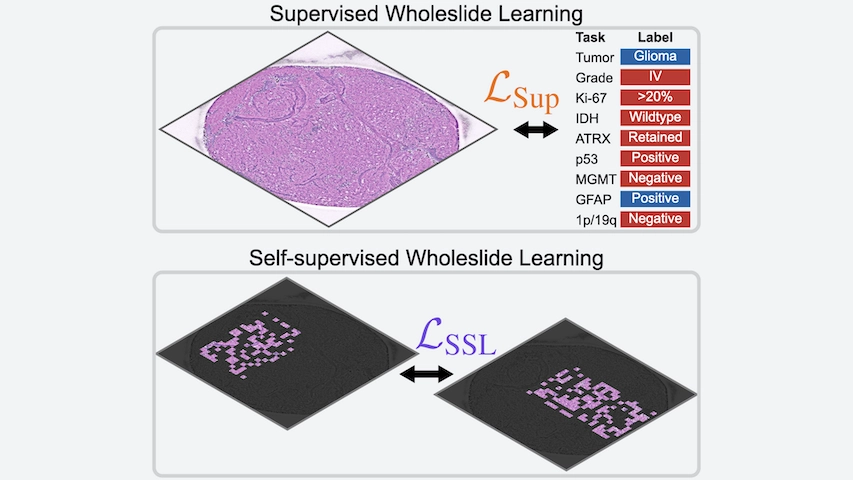
ARXIV · 2024This study introduces Slide Pre-trained Transformers (SPT), a self-supervised framework for whole-slide representation learning that captures multiscale histologic structure to support downstream pathology tasks with limited manual annotation.
-
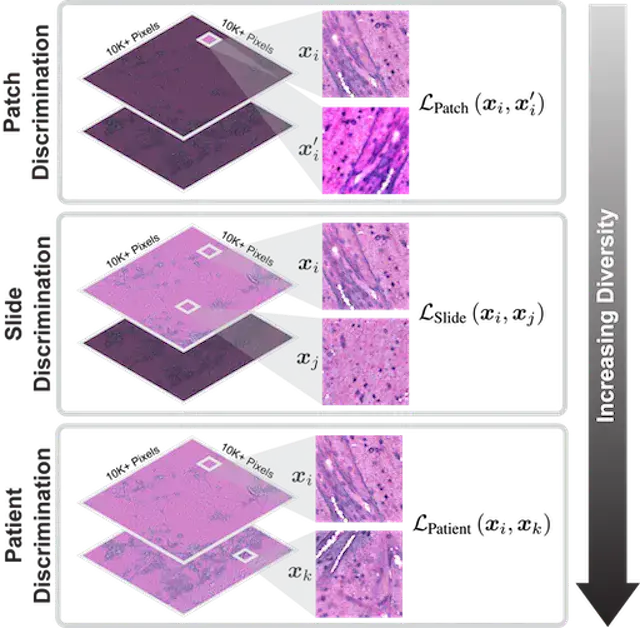
 COMPUTER VISION AND PATTERN RECOGNITION · 2023
COMPUTER VISION AND PATTERN RECOGNITION · 2023HiDisc is a self-supervised learning method that leverages the inherent patient-slide-patch hierarchy of biomedical microscopy to learn stronger visual representations without explicit negative mining.
-
 NEURIPS DATASETS & BENCHMARKS · 2022
NEURIPS DATASETS & BENCHMARKS · 2022OpenSRH is the first public dataset of clinical stimulated Raman histology images from brain tumor patients, released alongside benchmarks to accelerate machine learning research for intraoperative brain tumor diagnosis.
-
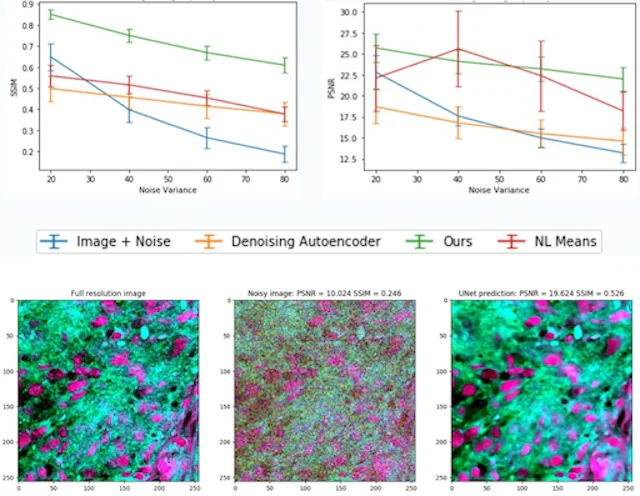
MLHC · 2020
This paper develops a weakly supervised denoising approach for stimulated Raman histology, improving image quality in label-free optical microscopy of human brain tumor specimens.