Todd Hollon
Intelligent Neuroimaging
A focus of the MLiNS lab is to develop clinically grounded artificial intelligence for neuroimaging by learning directly from routine health system data at scale. Our work has advanced from HLIP, which introduced hierarchical language-image pretraining for uncurated 3D MRI and CT studies, to ItemizedCLIP, which learns more complete and explainable visual representations from structured radiology supervision. Building on these foundations, we developed NeuroVFM, a generalist neuroimaging foundation model trained on millions of clinical MRI and CT volumes, and Prima, a health system-scale vision-language model for brain MRI designed for real-world diagnosis, triage, and clinical decision support. Together, this research program aims to create neuroimaging models that are accurate, interpretable, fair, and deployable across the full spectrum of neurologic disease, while establishing the health system itself as a powerful engine for medical AI discovery.
Figures
Related publications
-
 TRANSACTIONS ON MACHINE LEARNING RESEARCH · 2026
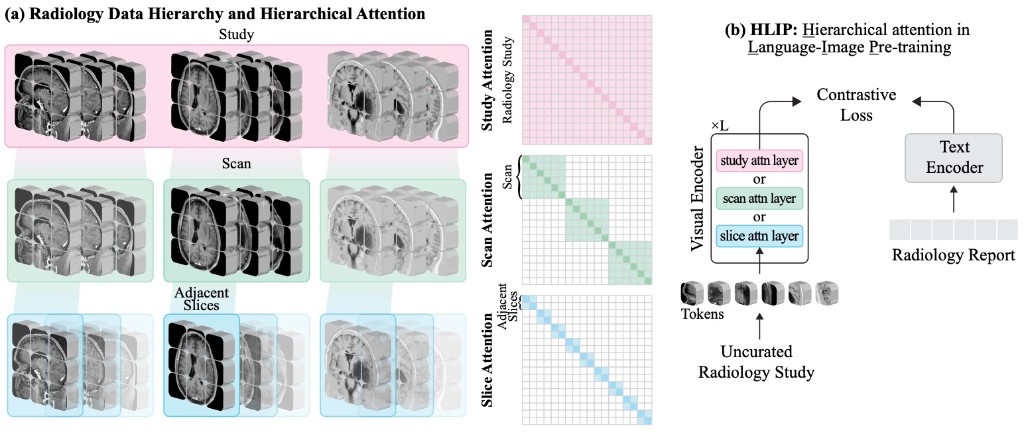
TRANSACTIONS ON MACHINE LEARNING RESEARCH · 2026Vision-language pre-training for volumetric MRI and CT is usually limited by radiologist-curated datasets. HLIP instead uses hierarchical attention over slice, scan, and study to pre-train on uncurated clinical data at scale, boosting performance on public brain MRI and head CT benchmarks.
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2026
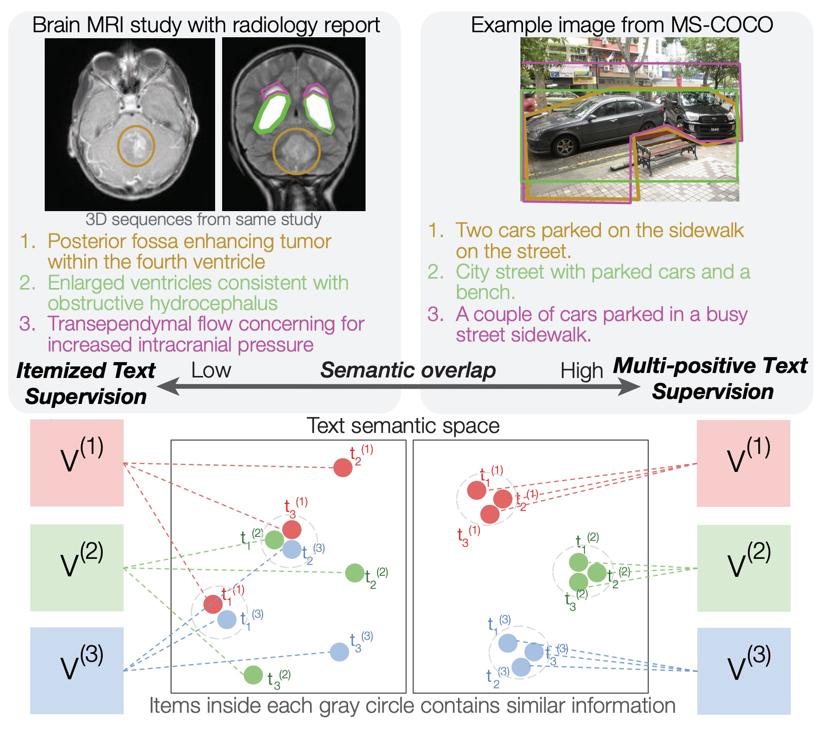
COMPUTER VISION AND PATTERN RECOGNITION · 2026Standard contrastive language-image pre-training can neglect objects in visual scenes. ItemizedCLIP forces models to learn and attend to all described items, resulting in better visual representations.
-
 arXiv · 2026
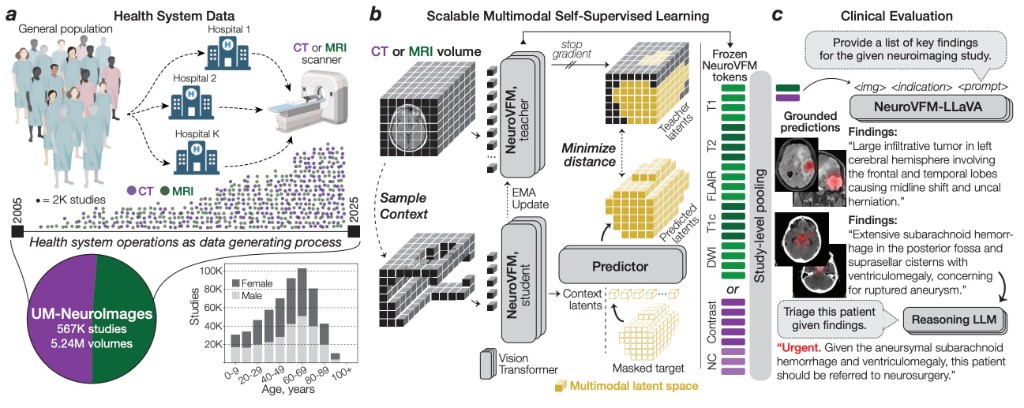
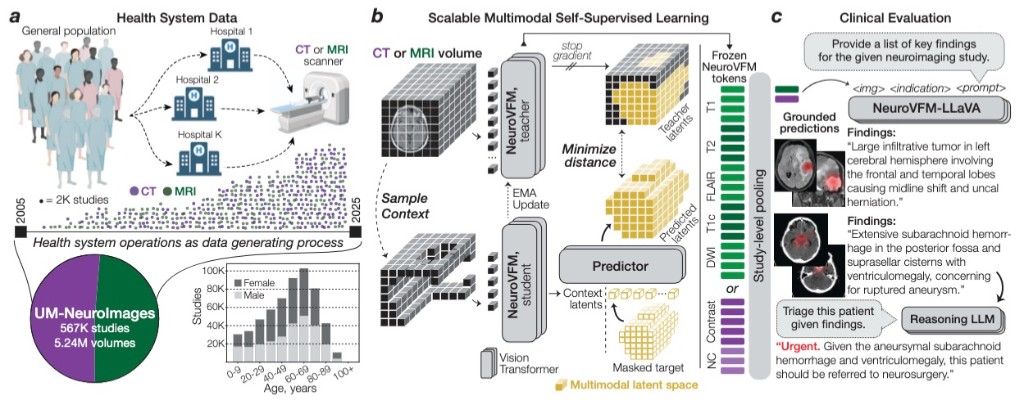
arXiv · 2026NeuroVFM is a visual foundation model trained on 5.24M clinical MRI and CT volumes via health system learning, a paradigm that leverages uncurated data from routine care. Using a scalable volumetric joint-embedding predictive architecture, it delivers state-of-the-art radiologic diagnosis and report generation with interpretable visual grounding, surpassing frontier models in accuracy, triage, and expert preference while reducing hallucinations.
-
 NATURE BIOMEDICAL ENGINEERING · 2026
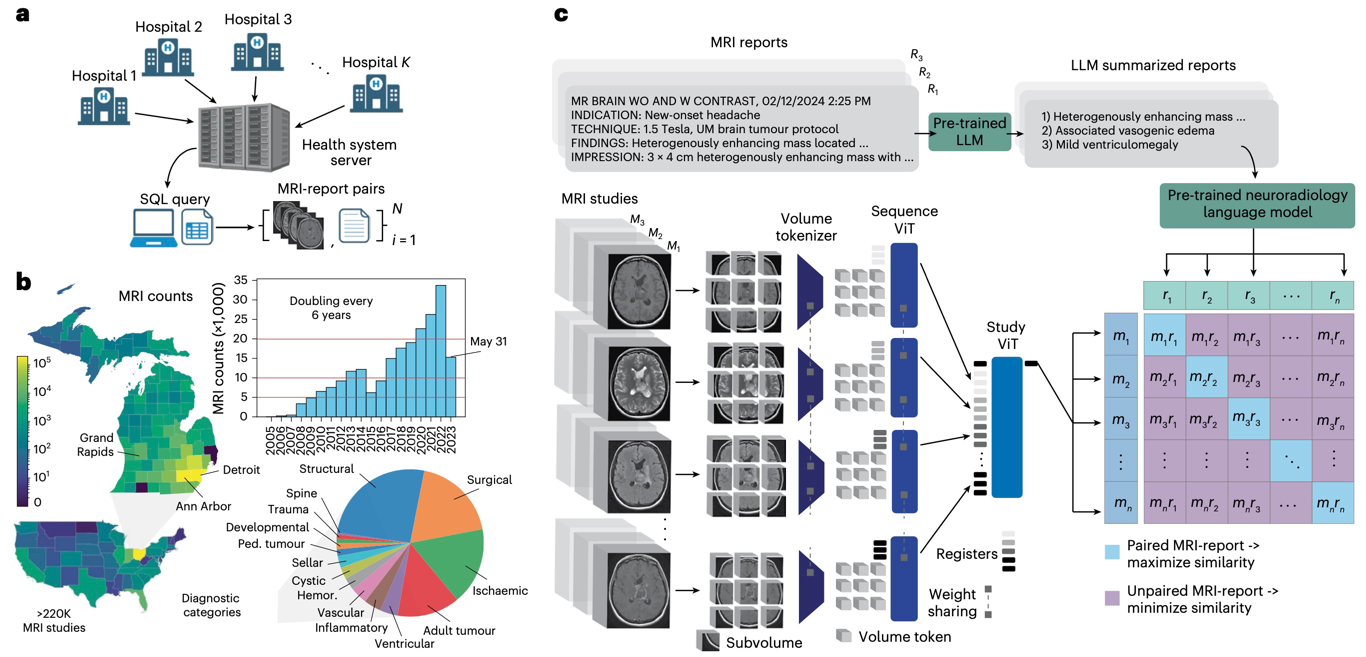
NATURE BIOMEDICAL ENGINEERING · 2026Introduces Prima, a visual foundation model for brain MRI trained at health-system scale for diagnosis, triage, and clinically grounded decision support.
-
 ARXIV · 2025
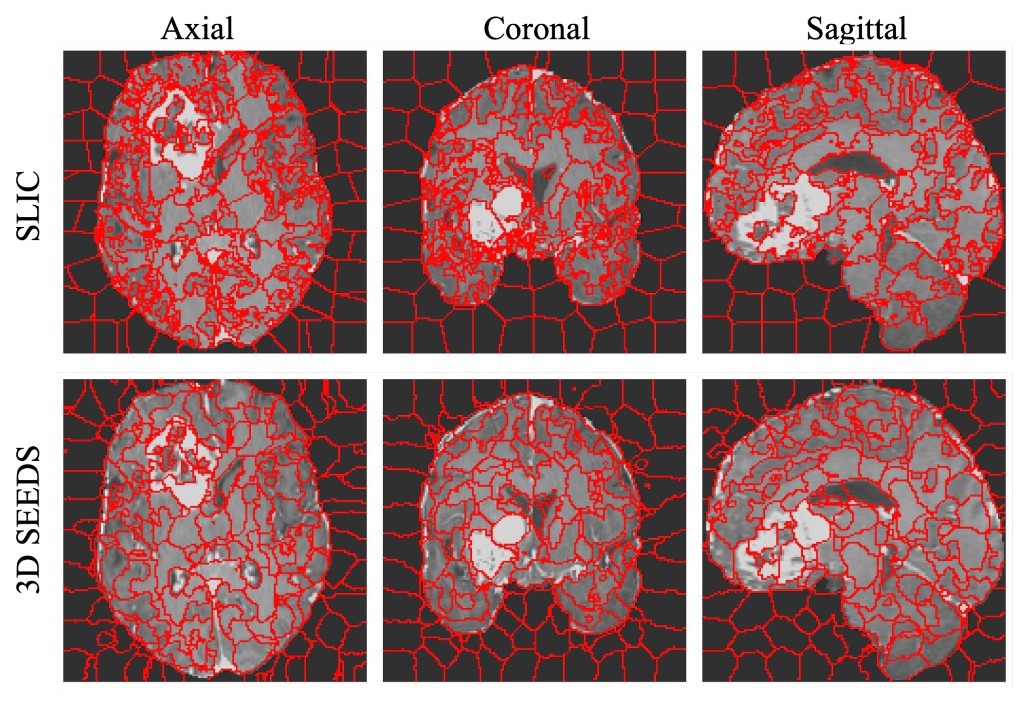
ARXIV · 2025This work extends the SEEDS superpixel algorithm from 2D to 3D volumes as 3D SEEDS, reporting substantially faster supervoxel generation and improved segmentation quality across diverse medical imaging tasks.
-
 NEUROSURGERY FOCUS · 2025
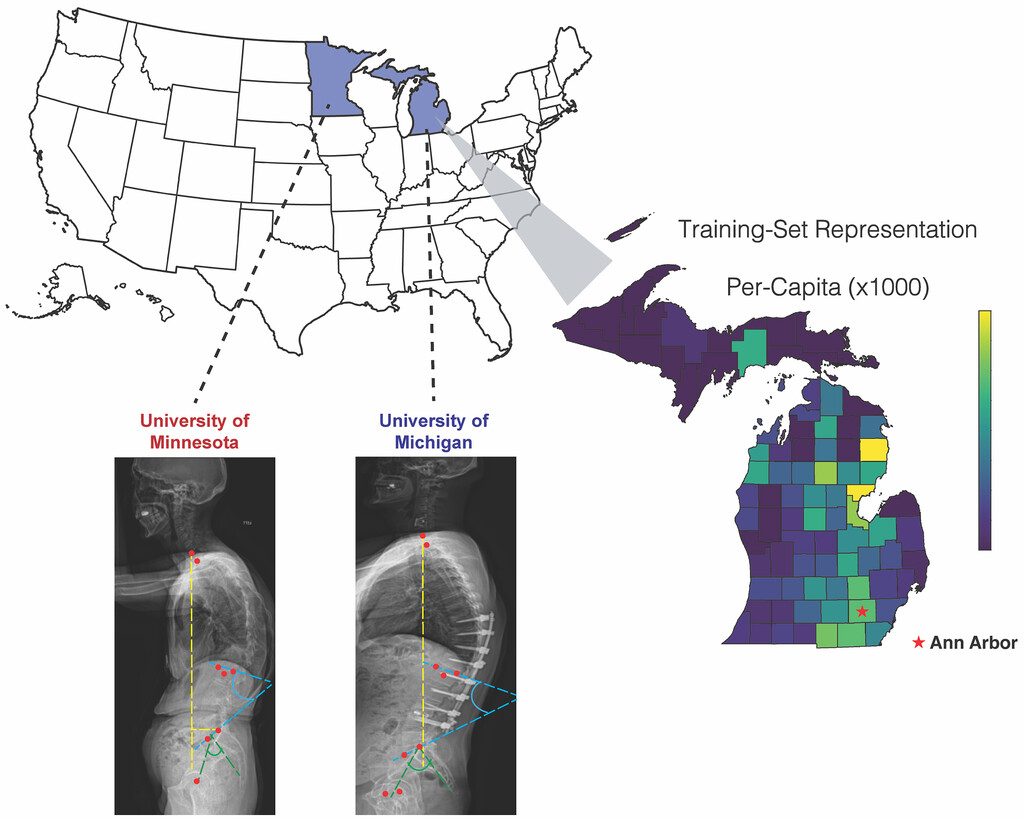
NEUROSURGERY FOCUS · 2025This external-cohort study validates SpinePose for automated spinopelvic parameter prediction from scoliosis radiographs, demonstrating generalizability across institutions.
-
 JOURNAL OF NEUROSURGERY SPINE · 2024
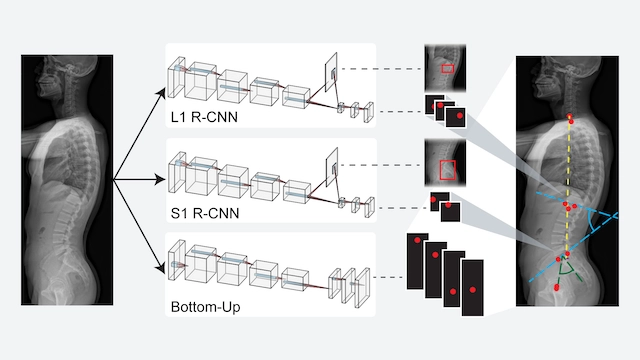
JOURNAL OF NEUROSURGERY SPINE · 2024This work introduces and validates an AI model for automated spinopelvic parameter estimation from imaging, aiming to improve speed and consistency in preoperative spinal alignment assessment.
-
PITUITARY · 2022
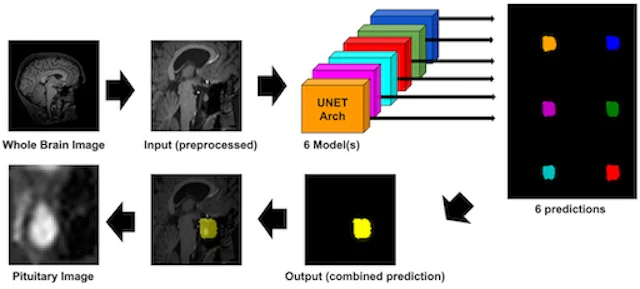
This study builds a new pituitary imaging resource by combining open-source scans with deep volumetric segmentation, enabling larger and more standardized datasets for pituitary AI research.