Todd Hollon
Intelligent Histology
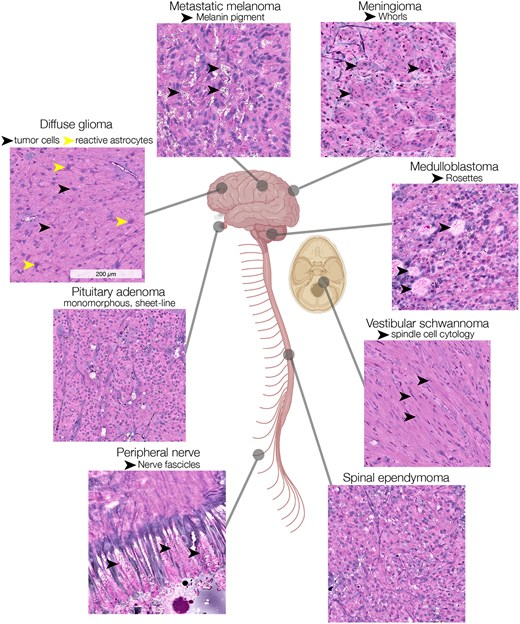
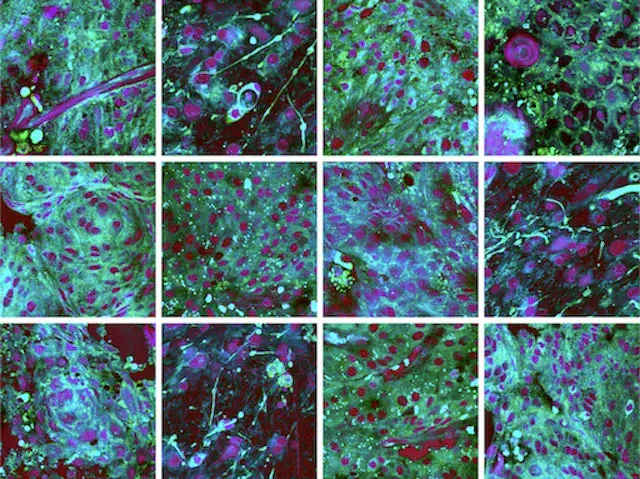
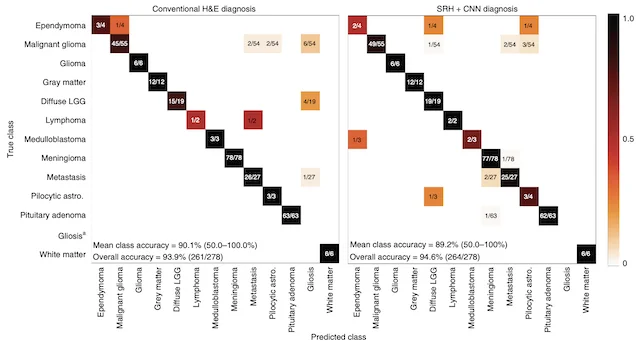
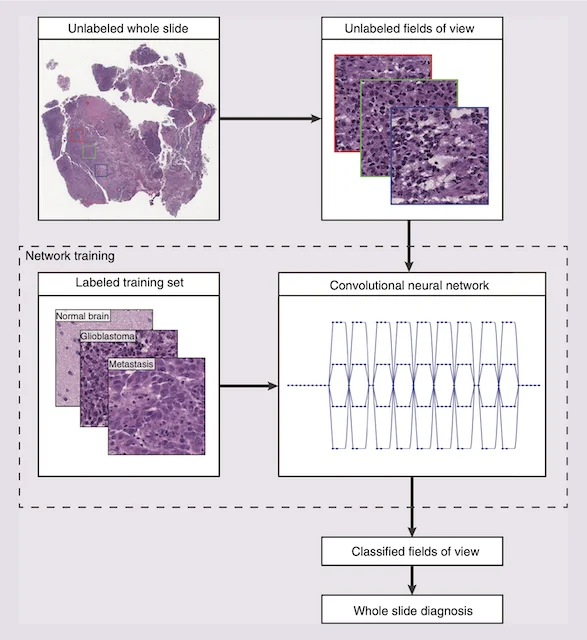
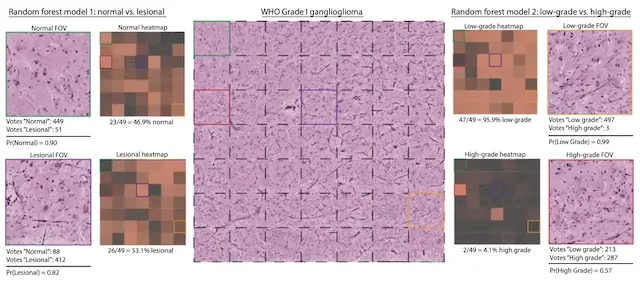
A major focus of the MLiNS lab is to combine stimulated Raman histology (SRH), a rapid label-free, optical imaging method, with deep learning and computer vision techniques to discover the molecular, cellular, and microanatomic features of skull base and malignant brain tumors. We are using SRH in our operating rooms to improve the speed and accuracy of brain tumor diagnosis. Our group has focused on deep learning-based computer vision methods for automated image interpretation, intraoperative diagnosis, and tumor margin delineation. Our work culminated in a multicenter, prospective, clinical trial, which demonstrated that AI interpretation of SRH images was equivalent in diagnostic accuracy to pathologist interpretation of conventional histology. We were able to show, for the first time, that a deep neural network is able to learn recognizable and interpretable histologic image features (e.g. tumor cellularity, nuclear morphology, infiltrative growth pattern, etc) in order to make a diagnosis. Our future work is directed at going beyond human-level interpretation towards identifying molecular/genetic features, single-cell classification, and predicting patient prognosis.
Figures
Related publications
-
 NEURO-ONCOLOGY ADVANCES · 2026
NEURO-ONCOLOGY ADVANCES · 2026This review synthesizes advances in intelligent histology for neurosurgery, highlighting how stimulated Raman histology and modern machine learning are converging to support faster, more precise intraoperative decision-making.
-
 STAR PROTOCOLS · 2025
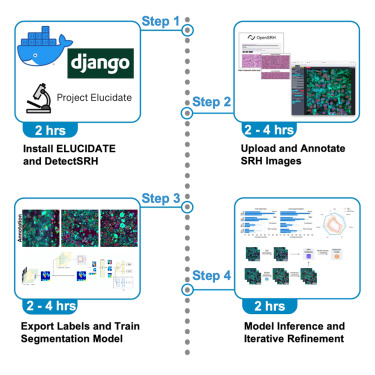
STAR PROTOCOLS · 2025This protocol provides a reproducible workflow for single-cell annotation and segmentation, called Elucidate, in stimulated Raman histology, enabling scalable dataset curation and model training for computational pathology studies.
-
 NATURE · 2025
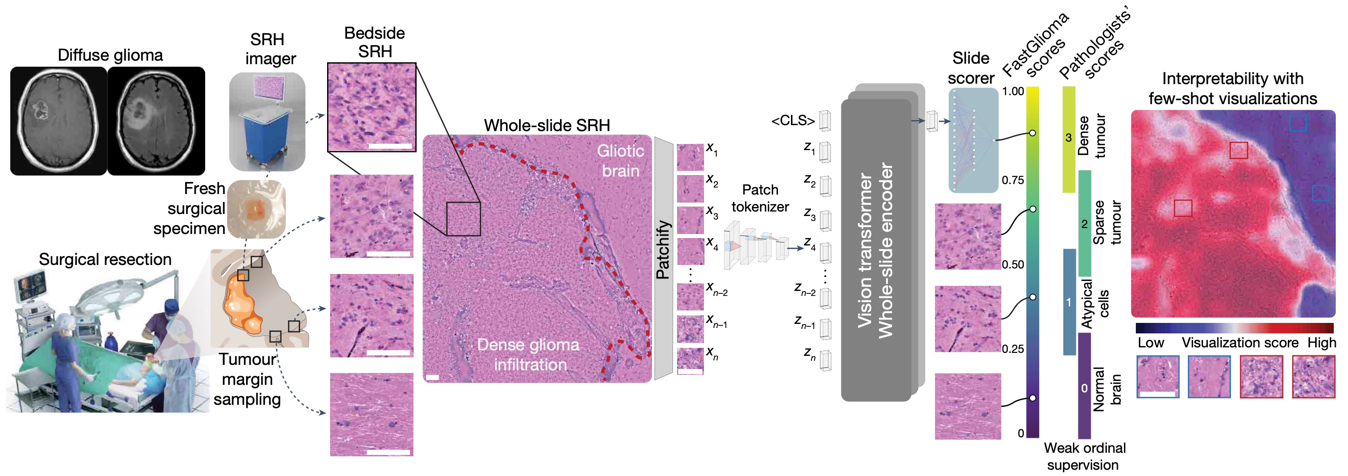
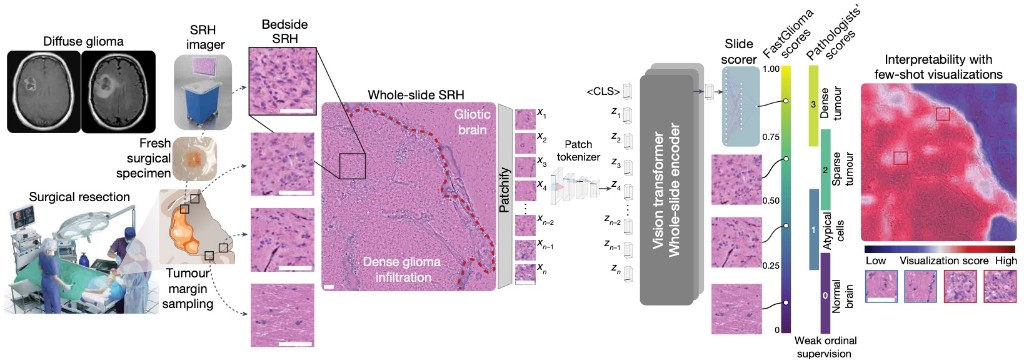
NATURE · 2025FastGlioma is a computational pathology model for real-time detection of glioma infiltration at the surgical margin, outperforming the current standard of care.
-
 NATURE MEDICINE · 2023
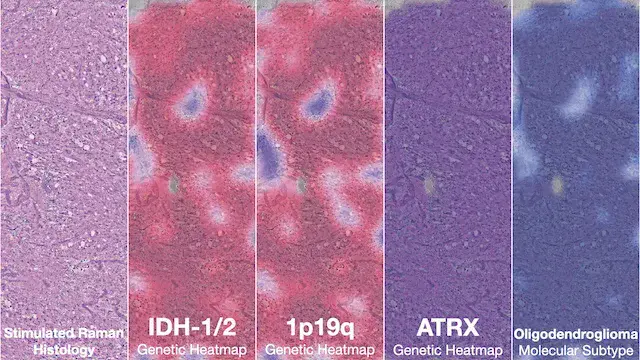
NATURE MEDICINE · 2023Diffuse gliomas are classified using the molecular features. DeepGlioma predicts the molecular genetics of brain tumors within minutes of biopsy, in the operating room, to better inform surgical goals.
-
 COMPUTER VISION AND PATTERN RECOGNITION · 2023
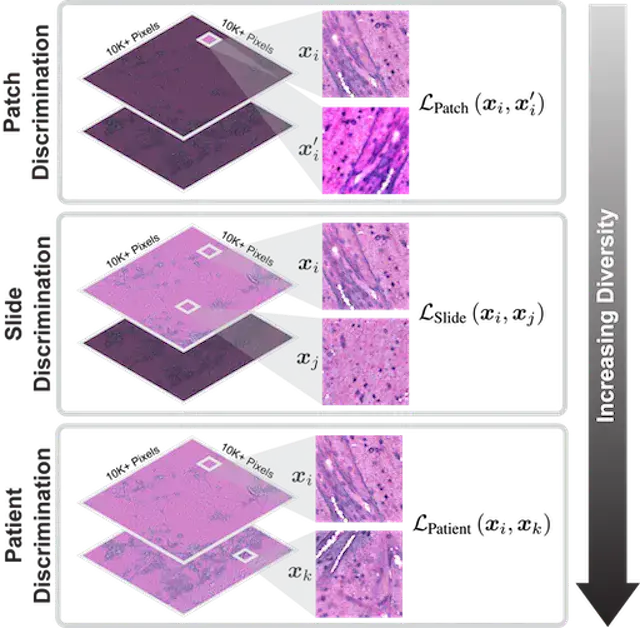
COMPUTER VISION AND PATTERN RECOGNITION · 2023HiDisc is a self-supervised learning method that leverages the inherent patient-slide-patch hierarchy of biomedical microscopy to learn stronger visual representations without explicit negative mining.
-
 NEURIPS DATASETS & BENCHMARKS · 2022
NEURIPS DATASETS & BENCHMARKS · 2022OpenSRH is the first public dataset of clinical stimulated Raman histology images from brain tumor patients, released alongside benchmarks to accelerate machine learning research for intraoperative brain tumor diagnosis.
-
 NATURE MEDICINE · 2020
NATURE MEDICINE · 2020A deep learning workflow combining stimulated Raman histology with convolutional neural networks delivers near real-time intraoperative brain tumor diagnosis, matching pathologist accuracy while compressing turnaround from ~30 minutes to under 150 seconds.
-
 NEURO-ONCOLOGY · 2021
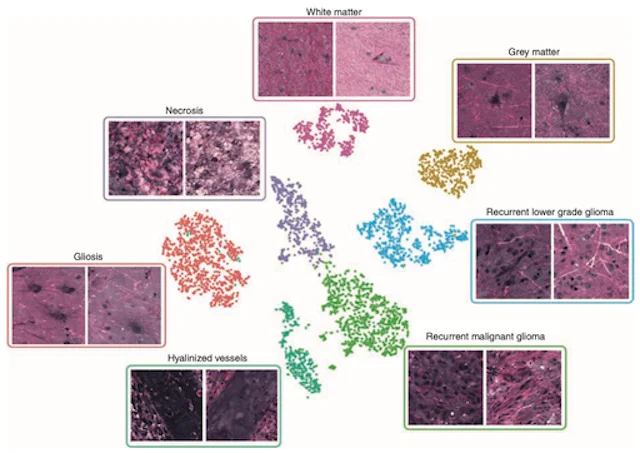
NEURO-ONCOLOGY · 2021This study demonstrates that stimulated Raman histology with deep neural networks can identify diffuse glioma recurrence intraoperatively, supporting rapid distinction between recurrent tumor and treatment-related changes.
-
CNS ONCOLOGY · 2020
This work presents an early machine learning framework for automated CNS tumor histologic diagnosis, showing how computational analysis can augment intraoperative pathology interpretation.
-
 CANCER RESEARCH · 2018
CANCER RESEARCH · 2018This paper extends stimulated Raman histology to pediatric neurosurgery, showing rapid, label-free intraoperative assessment of pediatric brain tumors to help guide operative management.
-
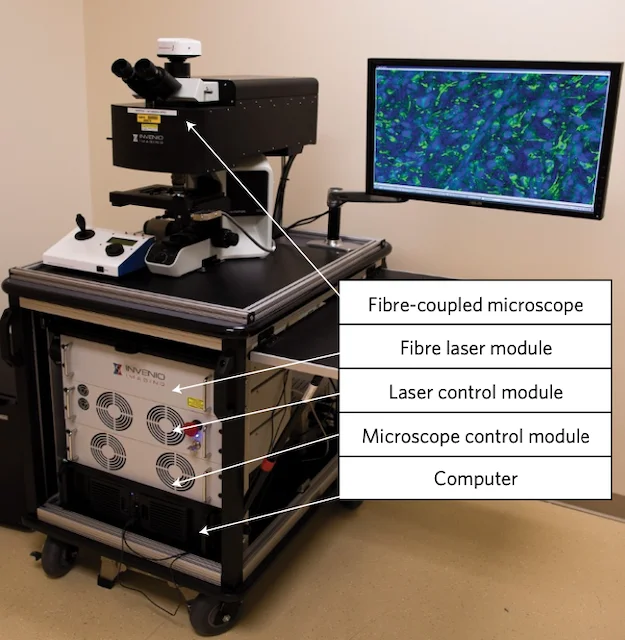
 NATURE BIOMEDICAL ENGINEERING · 2017
NATURE BIOMEDICAL ENGINEERING · 2017This study established stimulated Raman histology as a practical intraoperative microscopy method, enabling rapid, label-free visualization of fresh surgical tissue and laying the groundwork for real-time computational neuropathology.